New results from improved methods and models for SNP-based heritability and selection analyses
In a recently published article, professor Doug Speed from Center for Quantitative Genetics and Genomics (QGG), together with colleagues Anubhav Kaphle and David Balding (University of Melbourne), report their most up-to-date results concerning the genetic architectures of complex traits, and how their work with improved methods and models have corrected the original methods proposed more than a decade ago.
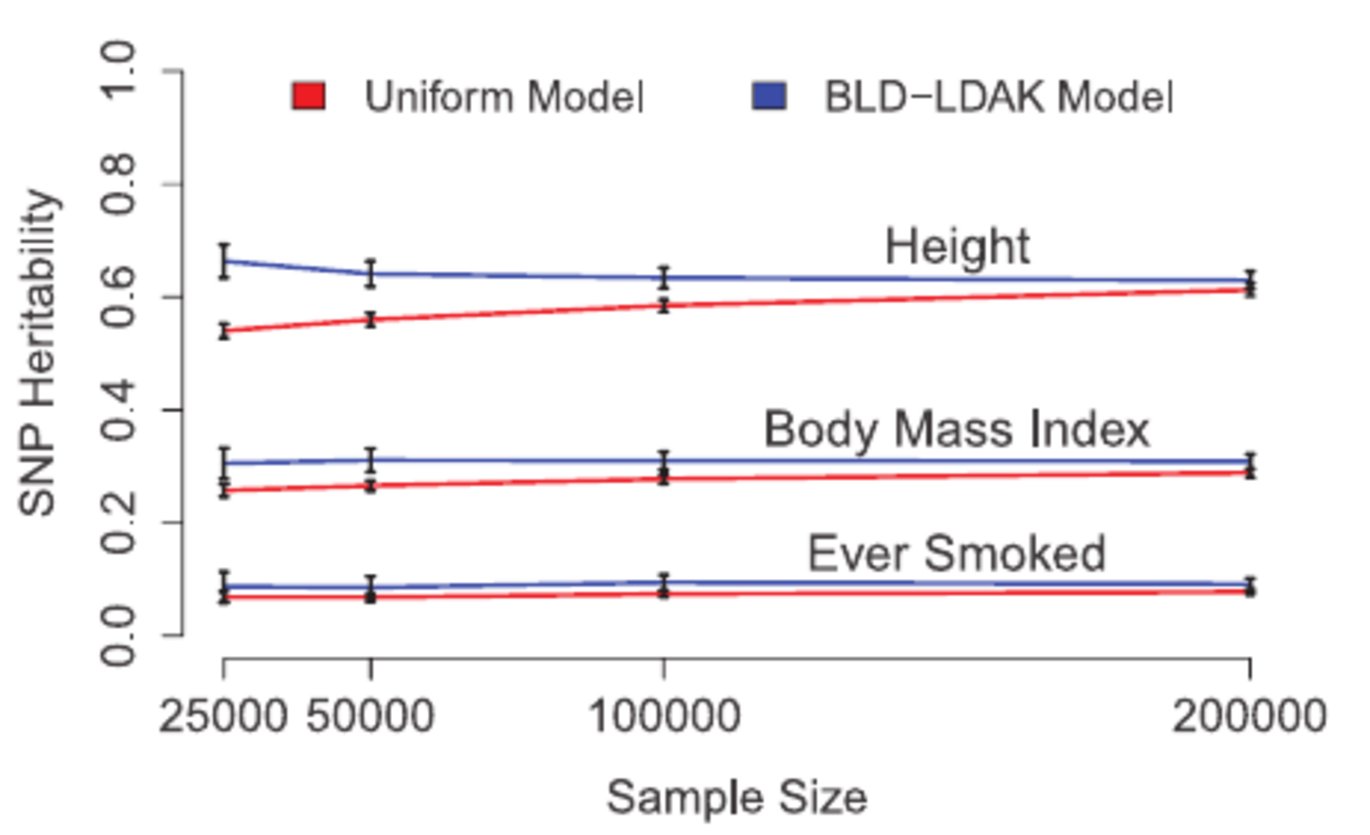
The article presents new results from analysing UK Biobank data. Specifically, it shows for a range of traits, how much of the heritability can be explained by different SNPs (and how this increases as we lower the minor allele threshold). It also reports updated estimates of heritability enrichments, showing the strong impact that conserved and coding regions have on a wide range of complex traits and diseases.
The article also provides a detailed tutorial on different methods for SNP-based heritability analyses. It explains the many ways these analyses enable us to better understand complex traits. For example, they can be used to estimate the total contribution of SNPs, to find the most important regions of the genome, and to measure the impact of selection pressures. The tutorial describes both methods that use individual-level data and those that use summary statistics.
Commenting on the article, Doug Speed says, "In this paper, we have attempted to explain a decade's worth of progress in the field of SNP-based heritability analysis. The original methods, first proposed in 2010, were adaptations of analyses designed for animal and plant breeding. Although these methods provided important insights into complex traits, they were fundamentally flawed because they were based on inappropriate modelling assumptions. Our new work first describes how new methods and models have been developed in order to address these errors. We then report the most up-to-date results concerning the genetic architectures of complex traits, including how these traits have been shaped by purifying selection".
Additional information | |
We strive to ensure that all our articles live up to the Danish universities' principles for good research communication (scroll down to find the English version on the web-site). Because of this the article will be supplemented with the following information: | |
Study type | Data analyses |
Funding | This research has been conducted using the UK Biobank Resource under Application Number 45918. Doug Speed is supported by the European Union Horizon 2020 Research and Innovation Programme under the Marie Skłodowska-Curie grant 754513, by Aarhus University Research Foundation (AUFF), by the Independent Research Fund Denmark under Project no. 702500094B, and by a Lundbeck Foundation Experiment Grant. This research was partially supported by grant DP190103188 from the Australian Research Council to David Balding and Doug Speed. |
Collaborators | Anubhav Kaphle and David Balding School of Mathematics and Statistics, University of Melbourne, Melbourne, Australia. |
Conflicts of interest | The authors declared no conflict of interest. |
Read more
| SNP-based heritability and selection analyses: Improved models and new results Doug Speed, Anubhav Kaphle, David J. Balding. BioEssays. 2022. https://doi.org/10.1002/bies.202100170 |
Contact | Professor Doug Speed Mail.: doug@qgg.au.dk |
