New software tool enables more accurate estimation of human disease heritability
Professor Doug Speed from Center for Quantitative Genetics and Genomics (QGG) has developed TetraHer, a new software for estimating the heritability of diseases, that addresses many problems with existing methods.
The heritability of a phenotype* is the proportion of phenotypic variation due to genetic factors. Estimates of heritability are of great value in statistical genetics. For example, they inform whether there is value in performing a genetic study of a phenotype, and they provide an upper bound for the accuracy of genetic prediction. When the phenotype is a human disease, the heritability indicates the potential of personalized medicine, i.e., how well we can use an individual's DNA to predict whether they will develop the disease, or to determine the most suitable treatment.
The newly developed TetraHer software tool addresses many problems with existing methods. Firstly, most existing methods for estimating heritability are designed for continuous phenotypes; when applied to diseases, they tend to produce biased estimates, particularly for diseases that are rare or have moderate heritability. TetraHer avoids these biases by taking into account the binary nature of diseases (i.e., the fact that all individuals are either affected or unaffected). Secondly, TetraHer can accommodate covariates (i.e., non-genetic factors that also contribute towards the disease). This is especially important for human diseases where risk can vary substantially with factors such as age and sex, so these should be considered. Thirdly, TetraHer can allow for ascertainment, which occurs when the prevalence of a disease within a study differs from that in the population. Fourthly, TetraHer is fast, taking only seconds to analyze even datasets containing 100,000s of individuals.
Doug Speed explains:
‘- We have not only demonstrated the accuracy of TetraHer, but also used the method to estimate the heritability of 229 diseases based on ICD-10 codes, which is a standardized method of classifying diseases used across the world. We identified 107 diseases with significant heritability, which will be a useful resource for future genetic studies of complex traits.’
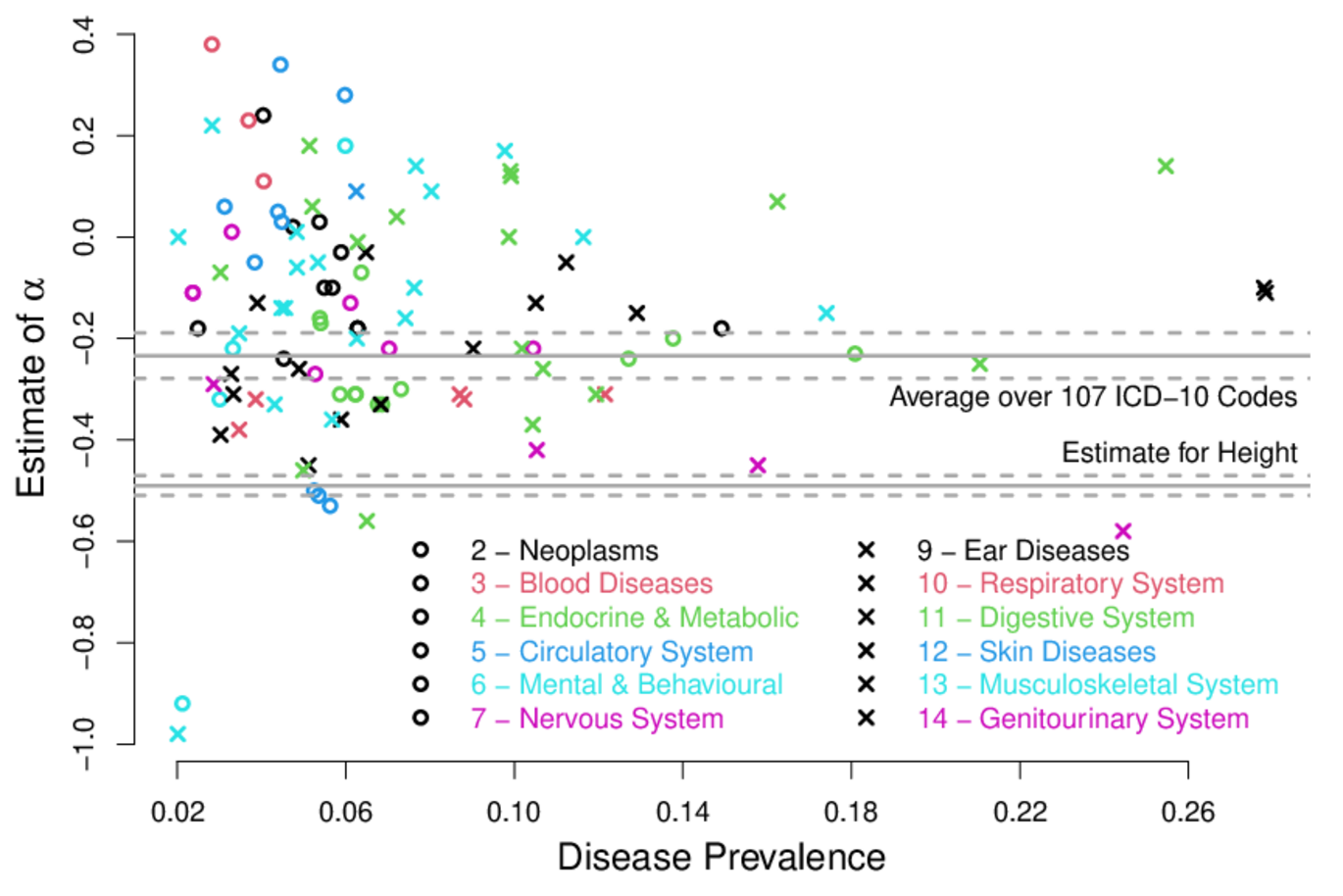
‘- For example, the figure above uses these diseases to investigate the relationship between the frequency and impact of risk variants. This relationship is an indicator of how much the disease influences fitness,’ he concludes.
Doug Speed has made TetraHer freely available within his software package LDAK. The article explaining the method has been published in the March issue of the American Journal of Human Genetics.
---
*Phenotype is an individual's appearance and/or characteristics, both visible (eg. gender, size, hair and eye colour) and physiological (eg. color vision, blood type). Source: Den store Danske.
