New and improved methods for predicting phenotypes for use in personalized medicine
Professor Doug Speed from Center for Quantitative Genetics and Genomics (QGG) and colleagues from the Bioinformatics Research Center (BiRC) and National Centre for Register-based Research (NCRR) at Aarhus University have recently developed improved methods for predicting complex traits. The work has this week been published in the journal Nature Communications.
There is currently great interest in being able to use an individual's genetic information to predict their phenotypes. This is especially important for personalized medicine, which aims to accurately predict which individuals will develop particular diseases or will benefit from particular medications.
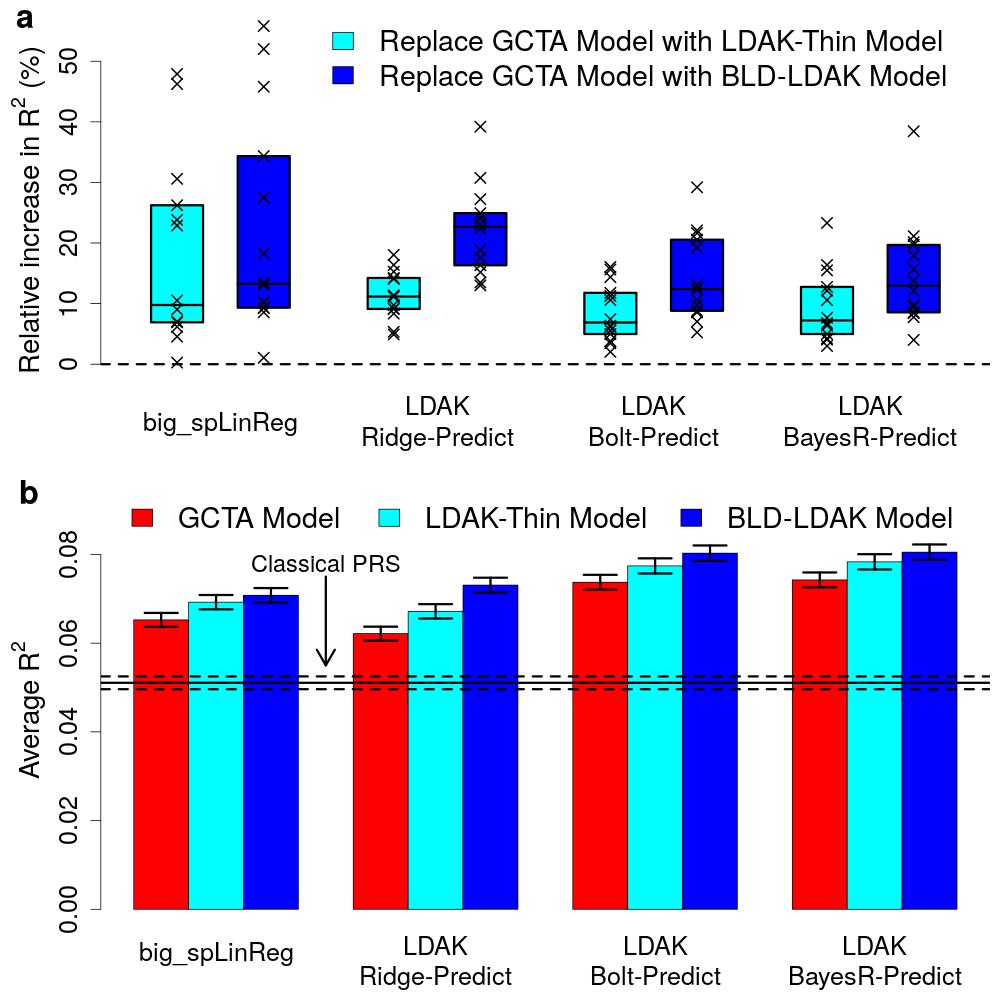
Doug Speed and his colleagues have observed that most existing prediction tools assume that each genetic variant is equally important. This assumption is sub-optimal, because recent work has shown that the importance of a variant depends on factors such as its frequency, local levels of linkage disequilibrium and functional annotations. Therefore, this new paper presents eight new prediction tools that allow for alternative assumptions, and shows that this enables substantially improved prediction across a wide range of traits.
Four of the new tools use individual-level data. The paper shows that the best of these, LDAK-Bolt-Predict, outperforms the existing tools Lasso, BLUP, Bolt-LMM and BayesR for all 14 phenotypes considered. The remaining four new tools use summary statistics. The paper shows that the best of these, LDAK-BayesR-SS, outperforms the existing tools lassosum, sBLUP, LDpred and SBayesR for 223 of the 225 phenotypes considered. On average, the new tools outperform the existing tools by 14% (sd 1), which is equivalent to increasing the sample size by about a quarter.
- ‘For personalized medicine to become a reality, we require models that can accurately predict an individual's phenotypes based on their genetic information. This work provides statistical tools for creating genetic prediction models that are substantially more accurate than existing tools’, Doug Speed explains, and continues:
- ‘This work will have immediate benefit, as it means we can now better identify individuals who have high risk of developing different diseases.’
You can read more about the new tools in the paper Improved genetic prediction of complex traits from individual-level data or summary statistics, and try out the new tools in the software packages LDAK and bigstatsr.
ITEM | CONTENT AND PURPOSE |
Type of study | We first developed new statistical tools, then compared their performance using data from UK Biobank, a population-based study of about 500,000 individuals recorded for a wide range of different phenotypes. We showed that our new tools outperformed existing tools for 223 out of 225 phenotypes considered. |
External collaborators | None |
External funding | Aarhus University Research Foundation, Lundbeck Foundation, EU Horizon 2020 Research and Innovation Programme, Independent Research Fund Denmark. |
Conflict of interest | The authors have declared no competing interest. |
Other | The article has been peer-reviewed. |
Link to the scientific article | You can read more about the new tools here: Zhang Q, Privé F, Vilhjálmsson B, Speed D. Improved genetic prediction of complex traits from individual-level data or summary statistics. Nature Communications. Try out the new tools in the software packages LDAK and bigstatsr. |
Contact information | Professor Doug Speed // doug@qgg.au.dk // Tel.: +44 77538 26477 |
