Researchers have developed an interactive mapping of gene expression and regulation in chickens
Researchers from the research consortium FarmGTEx, headed by tenure track assistant professor Lingzhao Fang from Center for Quantitative Genetics and Genomics (QGG), have developed the ChickenGTEx resource, a functional map of how variations affect gene expression in multiple chicken tissues. This can be used in the research on chicken genes – but farmers can, in the future, use it to breed chickens with specific traits.

What if farmers were in control of the genetic traits a chicken was born with? Researchers have recently developed a new web-portal, ChickenGTEx, where they have analyzed the genetic variations and identified regulatory variants in chickens.
In layman’s terms, this means that researchers have mapped out which DNA-variations affect how much a gene is expressed in different tissues. This makes it possible to see how tiny genetic differences can change a gene’s expression – without necessarily changing the gene itself. Here it becomes possible to track which of a chicken’s genes can result in unwanted traits such as slow growth, susceptibility to disease or problems with egg production.
What makes ChickenGTEx special is not only that it shows, which genes chickens possess, but also how tiny variations in the DNA affect how active the genes are in the different parts of the body.
The researchers have recently expanded the resource further to include egg-laying and sex-specific gene regulation. This gives the researchers insight into how genes’ ”on/off system” works in practice.
For instance, by using this resource, researchers have prioritized genes and tissues underlying a range of complex traits in chickens across egg laying and development stages.
By using the ChickenGTEx tool farmers can, in the future, select chickens with the most favorable genetic traits, so these can be bred – without having to do extensive laboratory tests to locate these chickens.
Infobox: On and off genes
Imagine DNA as the ‘recipe’ for a living creature. In the DNA strand lies, among other things, countless genes. These genes instruct the body in how a specific protein is produced.
Not all genes are active all the time. For example: a gene that helps cells produce energy is always on – because cells always need energy.
Other genes are only on under specific conditions – for example the genes that help in building muscle, which are active depending on the type of tissue, age, stress, environment or consumed nutrition.
There are also genes that are always inactive – in both humans and chickens. These are called ”pseudo-genes”. They have been rendered inactive through evolution and are therefore always off.
In this way there are on and off genes throughout the body – all depending on their function.
(From MedlinePlus Genetics)
ChickenGTEx: What is it?
ChickenGTEx is a digital portal developed by global researchers, led by tenure track assistant professor Lingzhao Fang from QGG at Aarhus University. It functions as a kind of interactive map of a chicken’s genes and DNA.
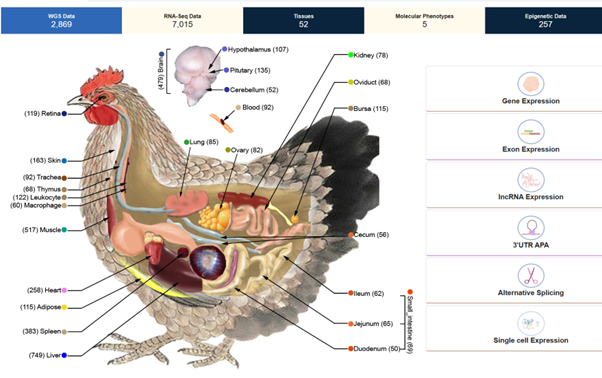
The map shows which genes can be found in the chicken as well as how the genes are turned on and off in the different tissues (heart, lungs, muscles etc.). This enables the researchers to see if the same DNA variation has different effects, for example in muscles and in the liver. This tissue-specific knowledge is crucial to understanding complex biological relationships.
Furthermore, the portal shows the DNA-variations in different chickens. It is these variations that affect chicken traits such as the colour of the feathers, growth or susceptibility to diseases.
The portal connects the genes with the traits in the chicken. This is done by comparing genetic data with documentation of specific traits. This enables the researchers to prioritize, which genetic variations are likely to play a key role in the development of specific traits. These traits could be qualities like egg production or growth.
In this way, the portal makes it possible to locate which genes are connected to the negative and positive traits, and it is based on this information that it becomes possible to select the chicken with the best genes for breeding.
Relevant for both farmers and researchers
The portal can prove ideal for farmers to select chickens with the best genes. Selective breeding of chickens with wanted traits can result in higher and better production of eggs and meat.
In reality it will most likely be breeding programs and genetic advisors that use this data, as it requires access to genetic analyses of the animals.
But the portal is equally important to researchers working with genetics. Researchers can use the portal to understand which genes control certain functions, when specific genes get turned on or off, tiny differences in DNA that affect gene expression and chicken traits. This also helps the researchers study evolution and how genes have developed over time.
The portal is a kind of interactive encyclopedia that can be used instead of researchers being forced to sequence DNA to analyze the genes.
“Future developments for ChickenGTEx will expand its scope to include more tissues and single-cell resolutions across various developmental stages. By integrating data from chickens under different environmental and external stimuli, we will create an increasingly robust and versatile functional resource. The ChickenGTEx resource will underpin the next generation of precision breeding strategies, leveraging the synergy between large-scale genomic language models and advanced biotechnologies, such as synthetic biology and CRISPR-based epigenetic editing.” says tenure track assistant professor Lingzhao Fang, leader of ChickenGTEx.
Chickens as a biological model
The mapping of the genes can also assist in the development of biomedicine. Chickens are often used as a kind of biological model in gene research, and the analysis of how the genes are turned on and off can also be used to better understand human genes.
By understanding these mechanisms in chickens, it is possible to learn more about how regulatory DNA functions in general. This knowledge can be transferred to studies in subjects such as cancer, metabolic diseases or disturbances in the immune system.
This ’atlas’ of chickens’ DNA enables researchers to compare gene regulation between species and understand common biological processes.
ChickenGTEx is an important resource within both the fields of farming and gene research, as it can, in the future, contribute to the biological knowledge that is the basis for the development of new medicine. This encyclopedia helps researchers understand the chickens’ DNA and which biological processes are in play. The overall goal of ChickenGTEx, as part of FarmGTEx project, is to generate a comprehensive map of regulatory effects on all genetic variants in global chicken genome. This is also why the researchers will keep expanding the ChickenGTEx resource in the future, to cover more tissues, cells, development stages, and breeds.
| Additional information | |
| We strive to ensure that all our articles live up to the Danish universities' principles for good research communication (scroll down to find the English version on the web-site). Because of this the article will be supplemented with the following information: | |
Study type
| Genetic analysis, database description |
Funding
| National Key Research and Development Program of China (2024YFF1000100). Agricultural Science and Technology Innovation Program (ASTIP-IAS-04-2). National Natural Science Foundation of China (T2425005 and 32030021). International Partnership Program of the Chinese Academy of Sciences (153F11KYSB20160008). Open Biodiversity and Health Big Data Programme of IUBS. Chinese Academy of Agricultural Sciences. |
Collaborators
| Yali Hou (Chinese Academy of Agricultural Sciences). Dong Zou (China National Center for Biomedicine). Qin Chu (Beijing Academy of Agriculture and Forestry Sciences). Ruizhen Wang (Academy of Chinese Medical Sciences). Dailu Guan (University of California). Wannian Wang (Shanxi Agricultural University). Xiao Feng (China Agricultural University). Xin Li (Chinese Academy of Agricultural Sciences). Xiaoning Zhu (China Agricultural University). Zhonghao Bai (China Agricultural University). Yahui Gao (South China Agricultural University). Hongwei Yin (Chinese Academy of Agricultural Sciences). Tianyi Xu (Beijing Institute of Genomics). Zhixiang Yuan (Beijing Institute of Genomics). Xiaoxiang Hu (China Agricultural University). Ning Yang (China Agricultural University). Huaijun Zhou (University of California). Lingzhao Fang (QGG, Aarhus University). Zhang Zhang (Chinese Academy of Sciences). |
| Conflicts of interest | The authors declare no conflict of interest |
| Read more | ChickenGTEx: ChickenGTEx portal: a pan-tissue catalogue of regulatory variants shaping transcriptomic and phenotypic diversity | Nucleic Acids Research | Oxford Academic |
| Contact | Tenure track assistant professor Lingzhao Fang |
